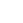
Professor Mike Thompson is a co-author of a paper in Science that expands the space of protein geometries by computational design. Protein design typically selects a protein topology and then identifies the geometries (secondary-structure lengths and orientations) that give the most stable structures. A challenge for this approach is that functional sites in natural proteins often adopt nonideal geometries. In this article, Pan et al. addressed this issue by exploring the diversity of geometries that can be sampled by a given topology. They developed a computational method called LUCS that systematically samples geometric variation in loop-helix-loop elements and applied it to two different topologies. This method generated families of well-folded proteins that include structures with non-native geometries. The ability to tune protein geometry may enable the custom design of new functions.



